Research
New studentship offered with Professor Eperon
for the MIBTP doctoral training scheme.
Mechanisms of cold-dependent neuroprotection
Hypothermia is an important therapeutic tool for mitigating the neurological damage resulting from ischaemic brain injuries after cardiac arrest, stroke and other neurological insults. This benefit arises from increased expression of a cold-shock protein, RBM3, which is an RNA-binding protein that has effects on translation of selected mRNAs. Thus, there is considerable therapeutic interest in understanding how RBM3 is upregulated in the cold and whether it is possible to develop drugs to reproduce this effect.
How does cold increase expression of RBM3? Recent research has shown that the gene for RBM3 contains a poison exon. Inclusion of this exon in the spliced product, the mRNA, leads to its degradation. However, in cold conditions this exon is missed out, or ‘skipped’, leading to stabilisation of the mRNA and thus increasing the levels of protein expressed. This effect seems to depend on temperature-dependent binding of hnRNPH1 to the exon, which promotes skipping (Lin et al., 2023, EMBO J. 42: e113168).
The purposes of this project are to investigate the molecular mechanisms by which temperature affects the binding of hnRNPH1 and other proteins to the exon and to establish how hnRNPH1 mediates skipping when it binds. We have recently shown that we can recapitulate the temperature-dependent skipping in vitro, when synthetic pre-mRNA is added to a nuclear extract. This means that the process is amenable to molecular methods. We will use classical methods, such as cross-linking and affinity purification to identify RNA-bound proteins, SHAPE and other analyses to map the secondary structure of the RNA, and nuclease protection to monitor the binding of splicing factors. Mutagenesis of the proteins and RNA will be used to establish whether these biochemical properties are correlated with the exon skipping effect.
We have developed single molecule methods for establishing the numbers of proteins bound to the pre-mRNA (e.g., Cherny et al., 2010, EMBO J. 29, 2161-2172; Jobbins et al., 2022, EMBO J. 41: e107640). This is especially important for hnRNPH1, where there are multiple potential binding sites and the repression of the exon might be strongly dependent on the numbers bound. We can use the same methods to see whether hnRNPH1 binding excludes normal splicing factors or is correlated with the recruitment of co-repressors. Finally, single molecule FRET analyses will be done to understand whether repression of the exon at low temperatures affects the structure and dynamics of the exon and its flanking introns. These investigations may open the way to the development in future of new drugs that, for example, stabilise the interactions between specific proteins.
For more information and how to apply go to:
https://le.ac.uk/study/research-degrees/funded-opportunities/bbsrc-mibtp
Two new papers published for sLOLA.
Two new papers published for the sLOLA, available to view:
New program for stoichiometry analysis of
single-molecule fluorescence bleaching steps and single-molecule tracking.
FluoroTensor is a python application program for analysis of single-molecule fluorescence data. The software is designed to automate colocalization analysis of up to three colour channels with chromatic aberration correction and calculate time series fluorescence intensity traces, building up a dataset automatically from many raw data files. The program can then be used to analyse photobleaching of the fluorescent molecules to determine the stoichiometry of components is biological complexes using powerful artificial intelligence models with high accuracy, and fast compute time even on large datasets. The program also features an add-in for single-molecule tracking which automates tracking and mean-square displacement fitting of particle trajectories. The program uses a fitting error-based approach for cleaning up convolved diffusivity distributions of molecules with heterogenous Brownian motion and fits the distributions using Gaussian mixture modelling to precisely return the diffusivity of each component in the mixture. All data can be exported to preformatted Excel spreadsheets to facilitate easy further processing and visualization.
FluoroTensor v6.6.8r Full User Guide
FluoroTensor v6.6.8r Quick-Start Reference Guide
Some recent news from the SpliceSelect sLoLa team
Back in 2018, we published a paper in which we showed that splicing enhancers can stimulate splicing even if they are only connected to the rest of the pre-mRNA by a flexible but non-RNA linker, such as repeats of hexaethylene glycol or abasic RNA. This showed that the proteins bound to the enhancer must connect with the rest of the splicing apparatus via 3D-diffusion, because they would not be able to slide or build complexes along the non-RNA linker (Jobbins et al., 2018; Nuc.Acids Res. 46, 2145-2158).
We were stimulated to explore this further by creating linkers that might have different physical properties (bulk, flexibility, etc.) and would be more resistant to nucleases. Alex Axer, who won a Humboldt fellowship in 2021, has created a range of new analogues of abasic RNA (or poly-ribo/arabinose monophosphate) with different substitutions on the 2’OH position or rigid ring structures. He is currently testing the effects of these analogues on base-pairing and recognition by enzymes when they are introduced into DNA or RNA oligonucleotides, in collaboration with Andrea Taladriz Sender (all in Strathclyde).
Much of the research in Leicester involves single molecule methods for detecting and counting the proteins that regulate splicing that are bound to a particular molecule of pre-mRNA in nuclear extracts. This relies upon the expression of proteins tagged with a fluorophore, such as mEGFP or mCherry, in the cells from which the proteins are made. We add RNAs molecule tagged with a fluorescent dye to the extracts. TIRF microscopy is then used to identify the complexes containing the labelled proteins and, and we count the numbers of proteins bound by detecting the number of steps in which the fluorescence bleaches. This can be a slow and rather subjective process, but Max Wills has written some new software that uses convolutional neural networks to automatically detect complexes and count the bleaching steps. This has speeded up the process dramatically, and we hope that it will soon be available for wider dissemination. Sumera Tubasum and Vasileios Paschalis are using this in their research on SMN2 and Bcl-X splicing.
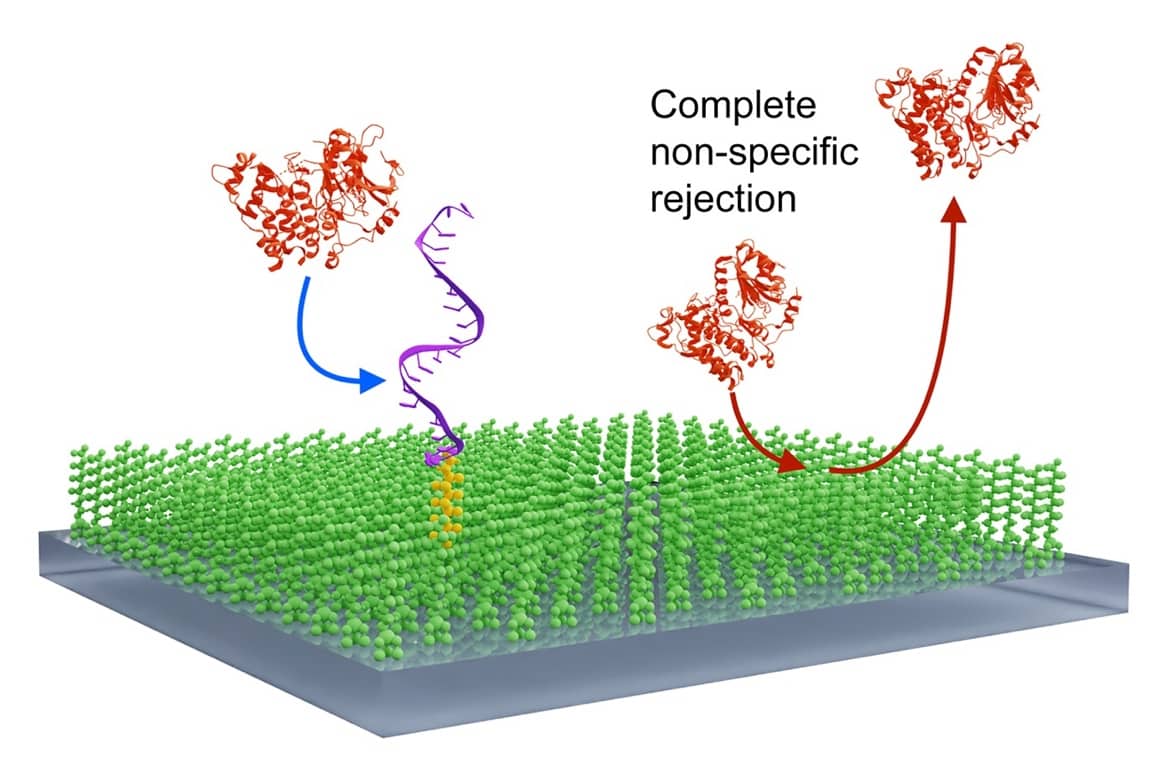
Most single molecule methods involve the use of surfaces on to which molecules are captured or deposited. These surfaces often cause headaches due to the high levels of non-specific binding. We have been testing fuorosilane surfaces as an alternative coating in the glass. Marina Santana Vega has been using the facilities in the School of Engineering in the University of Glasgow to create very high quality surfaces, and Carlos Bueno Alejo, in Leicester, has been testing them. They have remarkably low levels of adsorption: in iSCAMS, as many molecules rebound from the surface as collide with it in the first place (Bueno-Alejo et al., 2022; Appl. Mater. Interfaces 14, 49604-40616). Interestingly, macromolecules that are tethered to the surface via a fluorous ‘tail’ move around on the surface; co-migration could be a useful property for proving that two components have associated.
Seeing that proteins are bound together on an RNA molecule is not enough, though. We would like to know whether they interact, and how. For this, NMR is an ideal tool. However, working in functional nuclear extracts is challenging. The first question is whether the protein being studied has bound to the pre-mRNA, and where. Hesna Kara, in Leicester, has found that 19F substitution in specific sites in a pre-mRNA provides a good signal for detecting binding at that site, and moreover she has developed a much more sensitive signal that will really transform our ability to see binding at physiologically relevant protein concentrations. Alex Axer is exploring the incorporation of next-generation fluorinated nucleosides into RNA to enhance the sensitivity of these oligonucleotides for structural studies.
That’s it for now. In our next bulletin, we will provide more updates on the protein interactions being detected by Vasileios and Philippe de Gusmao Araujo, and the outcomes of the single molecule work being done by Sumera.